Areas of Interest
Research Interests
Structure, Assembly and Function of Cell Membranes, Lipids as Determinants of Membrane Protein Structure and Function, Role of Cardiolipin in Mitochondrial Organization and Function.
Structure, Assembly and Function of Cell Membrane Components
A better understanding of how lipid-protein interactions affect the structure and function of proteins is important for establishing the molecular basis for protein conformational disorders such as cystic fibrosis, Alzheimer’s disease, scabies, spinocerebellar ataxia type I, diabetes, sickle-cell anemia, and Nephrotic syndrome. Molecular genetic approaches are being used to construct strains of Escherichia coli in which membrane phospholipid composition can be regulated in a dose dependent and temporal manner to define the role of specific phospholipids in cell function. Through variation of phospholipid composition, the following roles for phospholipids have been defined: structure, topological organization and function of membrane proteins; function of the cell division and DNA replication machinery; organization of lipid domains in membranes; export of proteins across membranes. A fundamental objective in membrane biology is to understand and predict how protein sequences determine the number and orientation of transmembrane domains (TMDs). Through the use of strains of E. coli in which the synthesis of the major phospholipid, phosphatidylethanolamine (PE), can be regulated at steady state and temporally during the cell cycle, we have uncovered a central role of membrane phospholipid composition as a determinant of membrane protein topological organization. We have established that initial topological organization, existence of multiple topological conformers, and post-assembly topological re-organization of TMD orientation of membrane proteins are dictated by direct lipid-protein interactions. In more complex eukaryotic cells, changes in local lipid environment temporally or during intracellular vesicular tracking can change the organization and function of a membrane protein. Current projects focus on defining the molecular mechanism underlying the dynamic properties of membrane proteins in response to their lipid environment using genetic manipulation of proteins and lipid composition and biophysical and biochemical studies in living cells and reconstituted proteoliposomes.
Role of Cardiolipin in Supercomplex Formation
Cardiolipin is a major phospholipid found exclusively in the mitochondria of eukaryotic cells. We established that yeast mutants lacking cardiolipin fail to organize individual respiratory complexes into super complexes that make up the mitochondrial respirasome resulting in compromised respiratory function. Using structural (see SBIC Faculty section-Dowhan), genetic and biochemical approaches, our goal is to understand at the molecular level how cardiolipin organizes functional higher order molecular machines. Reduced cardiolipin levels are associated with several diseases as noted in the SBIC section.
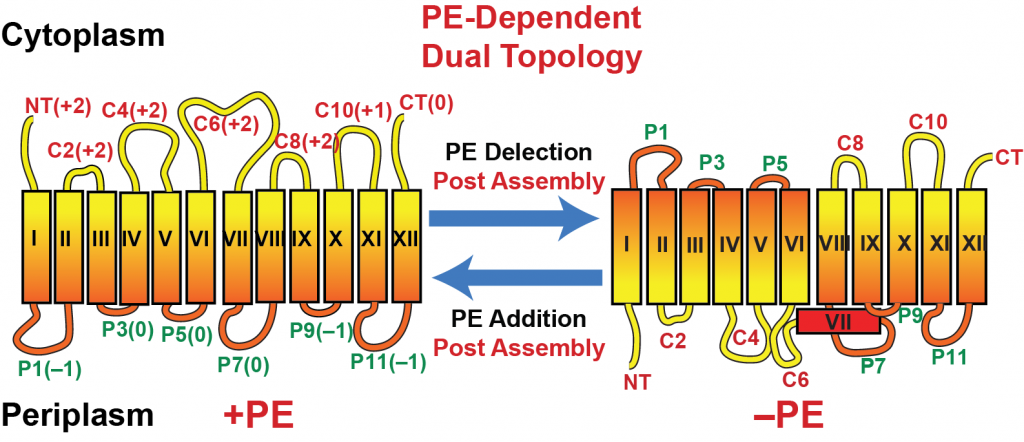
Topological Organization of Lactose Permease as a Function of Membrane Lipid Composition
Selected Publications
Vitrac H, MacLean DM, Jayaraman J, Bogdanov M, Dowhan W. Dynamic membrane protein topological switching upon changes in phospholipid environment. Proc Natl Acad Sci USA. 110, 13874-9 2015 PMCID 4653158
Ryabichko S, de Melo Ferreira V, Vitrac H, Kiyamova R, Dowhan W, Bogdanov M, Cardiolipin is required in vivo for the stability of bacterial translocon and optimal membrane protein translocation and insertion. Sci. Reports. 10, 6296 (2020) PMCID 7156725
Bogdanov M, Pyrshev K, Yesylevskyy S, Ryabichko S, Boiko V, Ivanchenko P, Kiyamova R, Guan Z, Ramseyer C, Dowhan W, Phospholipid distribution in the cytoplasmic membrane of Gram-negative bacteria is highly asymmetric, dynamic and cell shape dependent. Sci. Adv. 6, 6333 (2020) PMCID 726948
Hryc CF, Mallampalli VKPS, Bovshik EI, Azinas S, Fan G, Serysheva II, Sparagna GC, Baker ML, Mileykovskaya E, Dowhan W, Structural insights into cardiolipin replacement by phosphatidylglycerol in a cardiolipin-lacking yeast respiratory supercomplex using cryo-EM, Nat Comm 14, 2783 (2023)
Education and Training
AB
Chemistry, Princeton University
PhD
Biochemistry, University of California Berkeley
Postdoctoral Fellow
Biological Chemistry, Harvard Medical School
